Part 1 is here: statistic.
In November 2024 Nicolas Hulscher, Michael Cook, Raphael Stricker,
and Peter McCullough published a paper titled "Excess Cardiopulmonary Arrest and Mortality after
COVID-19 Vaccination in King County, Washington":
https://
The first author Nicolas Hulscher is the "foundation administrator" of McCullough Foundation, and he runs the Twitter account of McCullough Foundation.
Raphael Stricker is possibly the main person who popularized the hoax about Morgellons disease, and he was one of the two advisory board members of Morgellons Disease Foundation, which was founded by Mary Leitao who coined the term Morgellons disease. Stricker was prescribing ivermectin as a treatment for Morgellons before 2020. He was kicked out of academia in 1990 after he falsified data in a study about HIV, and afterwards he worked as an associate director of a penis enlargement clinic. In 2021 Stricker also authored a paper about hydroxychloroquine together with Peter McCullough and Harvey Risch, who are both members of the chief medical board of the Wellness Company. TWC's chief marketing officer used to be Christopher Alexander, whose bio says that he is "recognized as a leader in disinformation, misinformation, and counter-propaganda campaigns".
King County is the most populous county of the Seattle metropolitan
area. Hulscher and his coauthors compiled the data from yearly PDFs
published by the Emergency Medical Services Division of King County. For
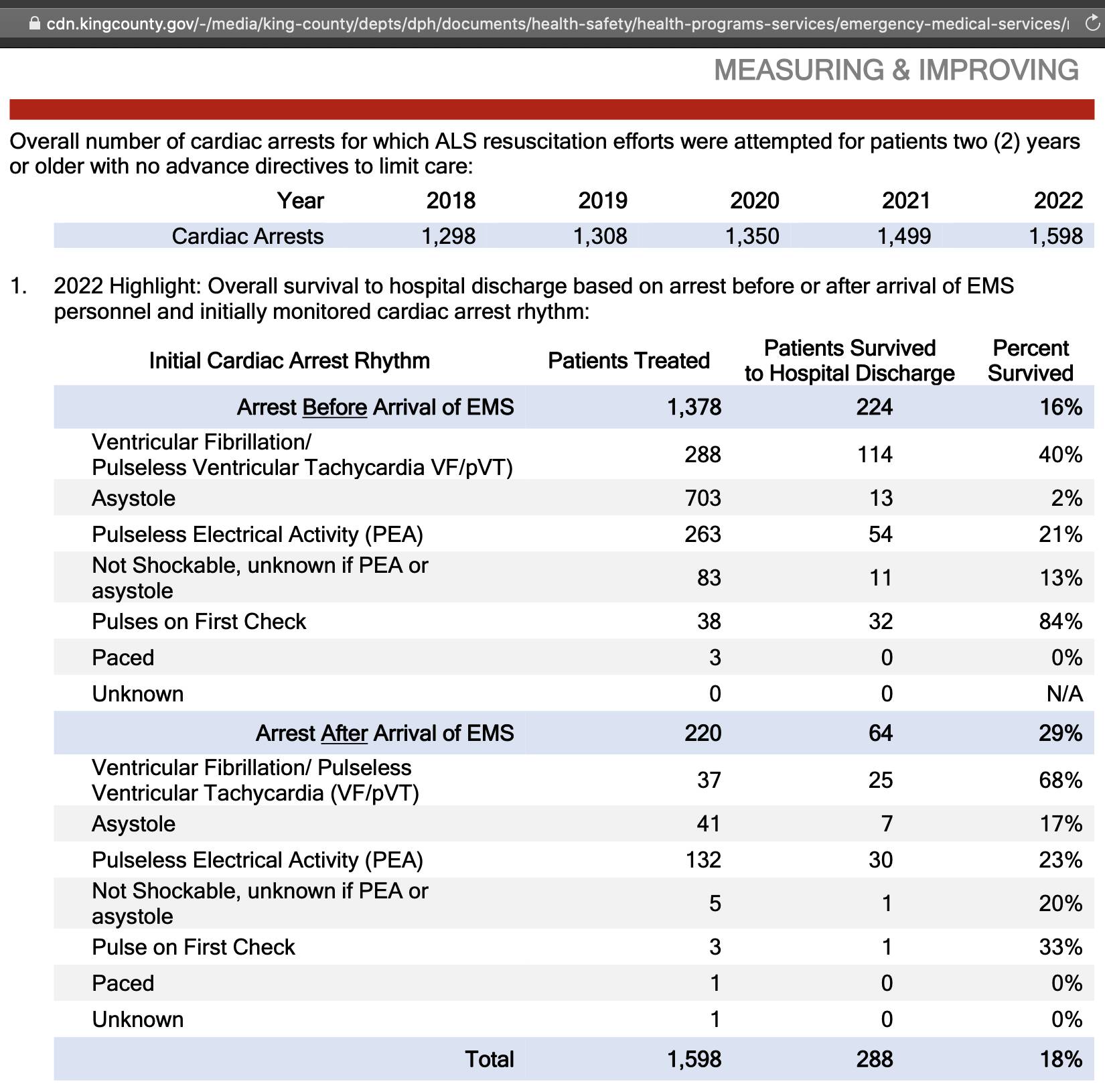
example the data for 2022 was shown in this table from the annual report
for 2023: [https://

The cardiac arrest cases that were included in the report were
defined as "cardiac arrests for which ALS
resuscitation efforts were attempted for patients two (2) years or older with no advance directives to
limit care". Hulscher seems to have calculated the number of
cardiac arrest deaths by multiplying the number of cases with the
rounded percentage of patients who didn't survive, which gave him a
figure of 1121 deaths in 2022 (from
1350*(). However he could've derived the
exact number of deaths by simply subtracting the number of survived
patients from the number of treated patients, so for example 1350 minus
234 would've given him the exact figure of 1116 deaths in 2020.
Hulscher also incorrectly entered the survival rate in 2021 as 18% even though it should've been 16%, but I didn't find other errors in his Table 1 when I compared it to the original PDF reports:
| Hulscher Table 1 | Original PDFs | |||||
|---|---|---|---|---|---|---|
| Year | Treated | Survived | Died | Treated | Survived | Died |
| 2005 | 1124 | 190 (17%) | 934 | |||
| 2006 | 993 | 174 (18%) | 819 | |||
| 2007 | 1035 | 191 (18%) | 844 | |||
| 2008 | 1046 | 199 (19%) | 847 | |||
| 2009 | 1072 | 206 (19%) | 866 | |||
| 2010 | 1069 | 218 (20%) | 851 | |||
| 2011 | 1047 | 223 (21%) | 824 | |||
| 2012 | 1134 | 252 (22%) | 882 | |||
| 2013 | 1135 | 235 (21%) | 900 | |||
| 2014 | 1246 | 267 (21%) | 979 | |||
| 2015 | 1114 | 20% | 891 | 1114 | 221 (20%) | 893 |
| 2016 | 1228 | 24% | 933 | 1228 | 288 (24%) | 940 |
| 2017 | 1215 | 21% | 960 | 1215 | 251 (21%) | 964 |
| 2018 | 1298 | 22% | 1012 | 1298 | 289 (22%) | 1009 |
| 2019 | 1308 | 19% | 1059 | 1308 | 253 (19%) | 1055 |
| 2020 | 1350 | 17% | 1121 | 1350 | 234 (17%) | 1116 |
| 2021 | 1499 | 18% | 1229 | 1499 | 242 (16%) | 1257 |
| 2022 | 1598 | 18% | 1310 | 1588 | 288 (18%) | 1300 |
| 2023 | 1697 * | 18% * | 1392 * | 1669 | 311 (19%) | 1358 |
Above I marked Hulscher's cells for 2023 with an asterisk because Hulscher et al. calculated them as a linear projection of the data for 2021 and 2022. In the real data for 2023 that was published in September 2024, the number of deaths was slightly lower than Hulscher's projected number of deaths for 2023:

I think Hulscher should've just omitted 2023 from his paper instead of including projected data for 2023, or in his plots he should've somehow visually differentiated the projected data from actual data.
On the website of Emergency Medical Services Division of King County, there's annual PDF reports going back to 2003, but the reports for 2003 and 2004 are missing the table which shows the number of treated cardiac arrest patients.
Hulscher misleadingly only included data from 2015 onwards in his paper. He used the years 2015-2020 as his baseline period, but in 2015-2020 the number of deaths fell unusually close to a line, which made the increase in deaths since 2021 seem fairly impressive in comparison. But during earlier years the deaths didn't follow such a neat trend, and for example there was a big drop in the number of deaths between 2014 and 2015. The data for 2015 was published in the report for 2016, but I didn't find any explanation for the drop in the report, I don't know if it there was a genuine decrease in deaths, or if there was some kind of a change in reporting practices, or if for example before 2015 the King County EMS used to serve a larger part of the Seattle metropolitan area than earlier.
There was also a fairly big drop in deaths between the years 2005 and
2006, but the PDF report for the year 2007 said that "the decrease in 2006 is due to a modification in case
definition": [https://

Hulscher used z-scores to indicate the level of excess deaths, but I think it would've been clearer to just express excess deaths as a percentage of the baseline number of deaths. His z-scores were exaggerated because the deaths fell unusually close to a line during his baseline period. Here when I simply added the year 2014 to my baseline period, my z-score for excess deaths in 2021 fell from about 13 to about 5:
d=data.table( year=2005: 2023) d$ dead=c( 934, 819, 844, 847, 866, 851, 824, 882, 900, 979, 893, 940, 964, 1009, 1055, 1116, 1257, 1300, 1358) d$ base1=d[ year% in% 2015: 2020, predict( lm( dead~ year), d)] d$ base2=d[ year% in% 2014: 2020, predict( lm( dead~ year), d)] d[ year==2021, dead- base1]/ d[ year% in% 2015: 2020, mean( abs( dead- base1))] # 13. d[24 year==2021, dead- base2]/ d[ year% in% 2014: 2020, mean( abs( dead- base2))] # 4. 662179
Hulscher started the x-axis of his plot from the year 2015, so you couldn't see how the long-term trend in the number of deaths in the EMS data appears to be curved upwards:

library(data. table); library( ggplot2) d=data. table( x=2005: 2023) d$ y=c( 934, 819, 844, 847, 866, 851, 824, 882, 900, 979, 893, 940, 964, 1009, 1055, 1116, 1257, 1300, 1358) d$ z=1; d$ proj=F base1=2015: 2020; base2=2006: 2019 p1=d[,.( x, y=d[ x% in% base1, predict( lm( y~ x), d)], z=2)][, proj: =! x% in% base1] p2=d[,.( x, y=d[ x% in% base2, predict( lm( y~ poly( x, 2)), d)], z=3)][, proj: =! x% in% base2] p=rbind( d, p1, p2) p[, z: =factor( z,, c( " Actual deaths", " 2015- 2020 linear trend", " 2006- 2019 quadratic trend"))] xstart=min( p$ x); xend=max( p$ x); ybreak=pretty( c( 0, p$ y), 7) ggplot( p, aes( x, y))+ geom_ point( aes( alpha=z, color=z), stroke=0, size=1. 2)+ geom_ line( data=p[ proj==F], aes( color=z), linewidth=. 4)+ geom_ line( aes( color=z), linewidth=. 4, linetype=" 11")+ labs( x=NULL, y=NULL, title=" Yearly cardiac arrest deaths in King County EMS data")+ scale_ x_ continuous( limits=c( xstart-. 5, xend+. 5), breaks=xstart: xend)+ scale_ y_ continuous( limits=range( ybreak), breaks=ybreak)+ scale_ color_ manual( values=c( " black", hsv( 22/ 36, 1,. 8), "# 00aa00"))+ scale_ alpha_ manual( values=c( 1, 0, 0))+ coord_ cartesian( clip=" off", expand=F)+ theme( axis. text=element_ text( size=11, color=" black"), axis. text. x=element_ text( angle=90, vjust=. 5, hjust=1), axis. ticks=element_ line( linewidth=. 4, color=" black"), axis. ticks. length=unit( 4, " pt"), axis. ticks. length. x=unit( 0, " pt"), legend. background=element_ blank(), legend. box. background=element_ rect( linewidth=. 4), legend. justification=c( 0, 1), legend. key=element_ blank(), legend. key. height=unit( 13, " pt"), legend. key. width=unit( 26, " pt"), legend. margin=margin( 4, 5, 4, 4), legend. position=c( 0, 1), legend. spacing. x=unit( 2,