Ethical Skeptic posted the plot below and wrote: [https://
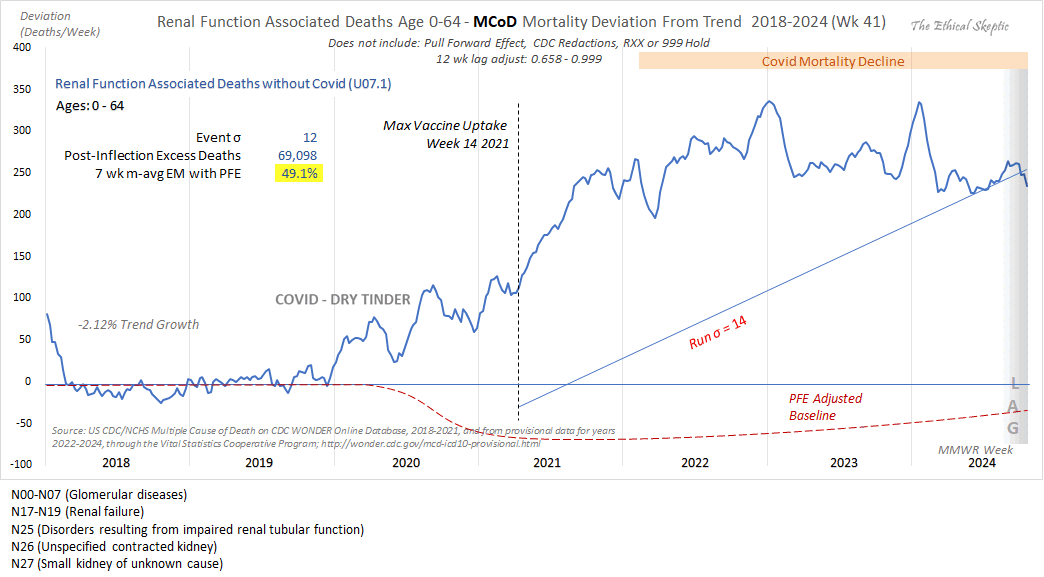
Since Kidney-related mortality was our #1 death excess this week, I took a look at its DFT chart.
While there is a Covid/Treatment effect in 2020. The overall arrival form still fits the VERY familiar Wk 14 2021 inflection/death taper.
This is the mRNA vaccine = 49% excess

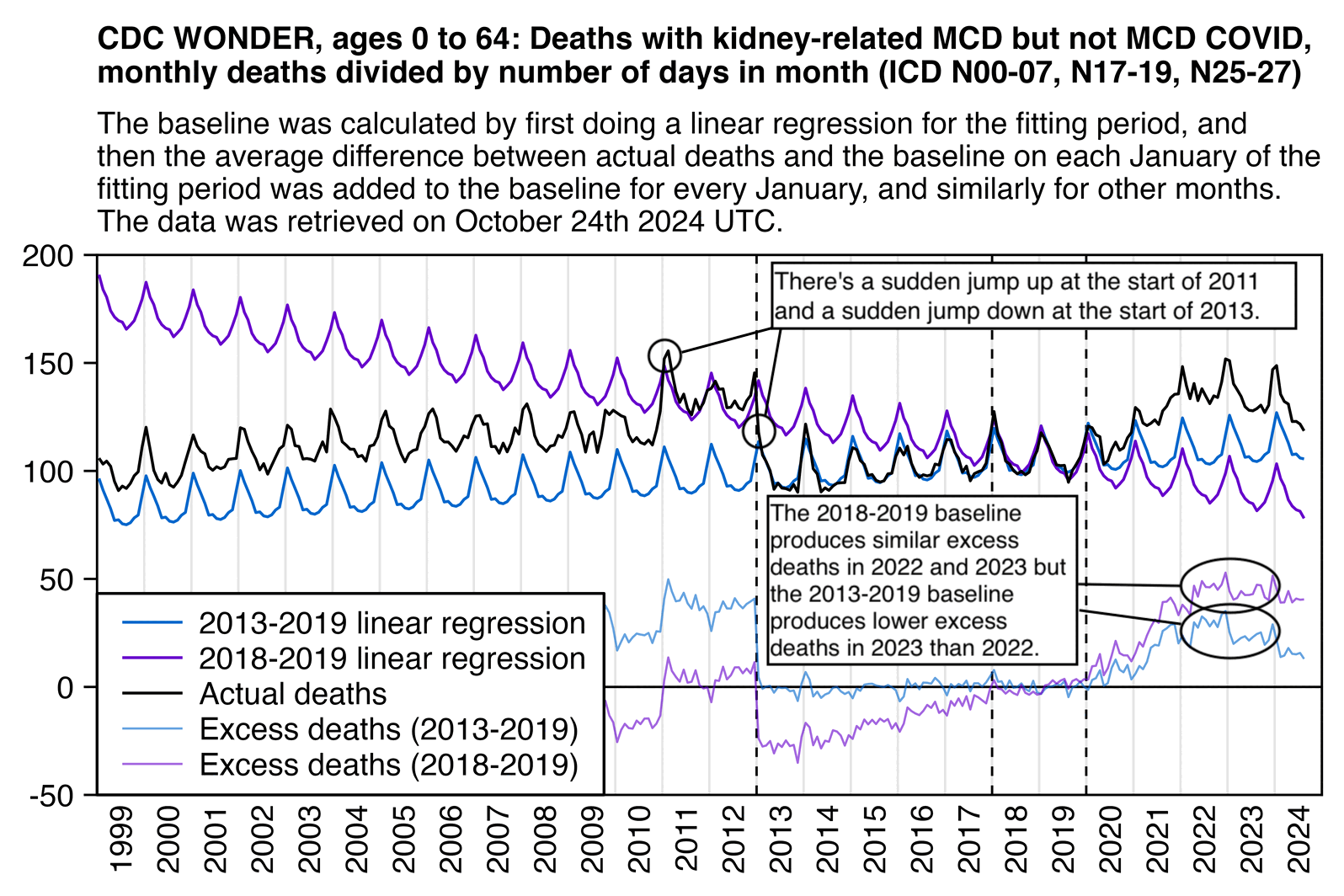
The reason why Ethical Skeptic got relatively high excess deaths in 2023 compared to 2022 might be if he fitted the baseline against data from only 2018 and 2019 as is demonstrated by my plot below. My plot also shows that the number of deaths under his ICD codes had a sudden jump up at the start of 2011 and a sudden jump down at the start of 2013, which may have been due to changes in coding practices. So the increase in deaths since 2020 might have similarly been due to changes to coding practices, because its magnitude is similar to the magnitude of the drop between 2012 and 2013:
library(data. table); library( ggplot2) t=fread( " http:// sars2. net/ f/ wonderkidney064. csv")[ date< =" 2024- 08"] t=merge( t[ cause==" MCD"], t[ cause==" MCD and COVID",.( date, and=dead)], all=T) t=t[,.( x=as. Date( paste0( date, "- 1")), y=dead- nafill( and,, 0), z=" Actual deaths")] t[, y: =y/ lubridate:: days_ in_ month( x)] a=rbind( copy( t)[, group: =2018], copy( t)[, group: =2013]) a[, month: =month( x)] a=merge( a, a[ year( x) < 2020& year( x) > =group,.( x=t$ x, base=predict( lm( y~ x), t)), group]) a=merge( a[ year( x) < 2020& year( x) > =group,.( monthly=mean( y- base)),.( month, group)], a) p=rbind( a[,.( x, y=base+ monthly, z=paste0( group, "- 2019 linear regression"))], t) p=rbind( p, a[,.( x, y=y-( base+ monthly), z=paste0( " Excess deaths (", group, "- 2019) "))]) p[, z: =factor( z, unique( z))] xstart=as. Date( " 1999- 1- 1"); xend=as. Date( " 2025- 1- 1") xbreak=seq( xstart, xend, " 6 month") xlab=c( rbind( " ", 1999: 2024), " ") ybreak=pretty( p$ y, 6)[- 1]; ystart=ybreak[ 1]; yend=max( ybreak) color=c( " black", hsv( 21/ 36, 1,. 8), hsv( 27/ 36, 1,. 8))[ c( 2, 3, 1, 2, 3)] ggplot( p, aes( x+ 15, y))+ geom_ vline( xintercept=seq( xstart, xend, " year"), color=" gray90", linewidth=. 25)+ geom_ vline( xintercept=as. Date( paste0( c( 2013, 2018, 2020), "- 1- 1")), linetype=2, linewidth=. 25)+ geom_ hline( yintercept=c( ystart, 0, yend), linewidth=. 25, lineend=" square")+ geom_ vline( xintercept=c( xstart, xend), linewidth=. 25, lineend=" square")+ geom_ line( aes( y=y, color=z, alpha=z, linewidth=z))+ labs( title=" CDC WONDER, ages 0 to 64: Deaths with kidney- related MCD but not MCD COVID,\ nmonthly deaths divided by number of days in month (ICD- 10 113 cause list\ n\ " Nephritis, nephrotic syndrome and nephrosis\ "; N00- 07, N17-