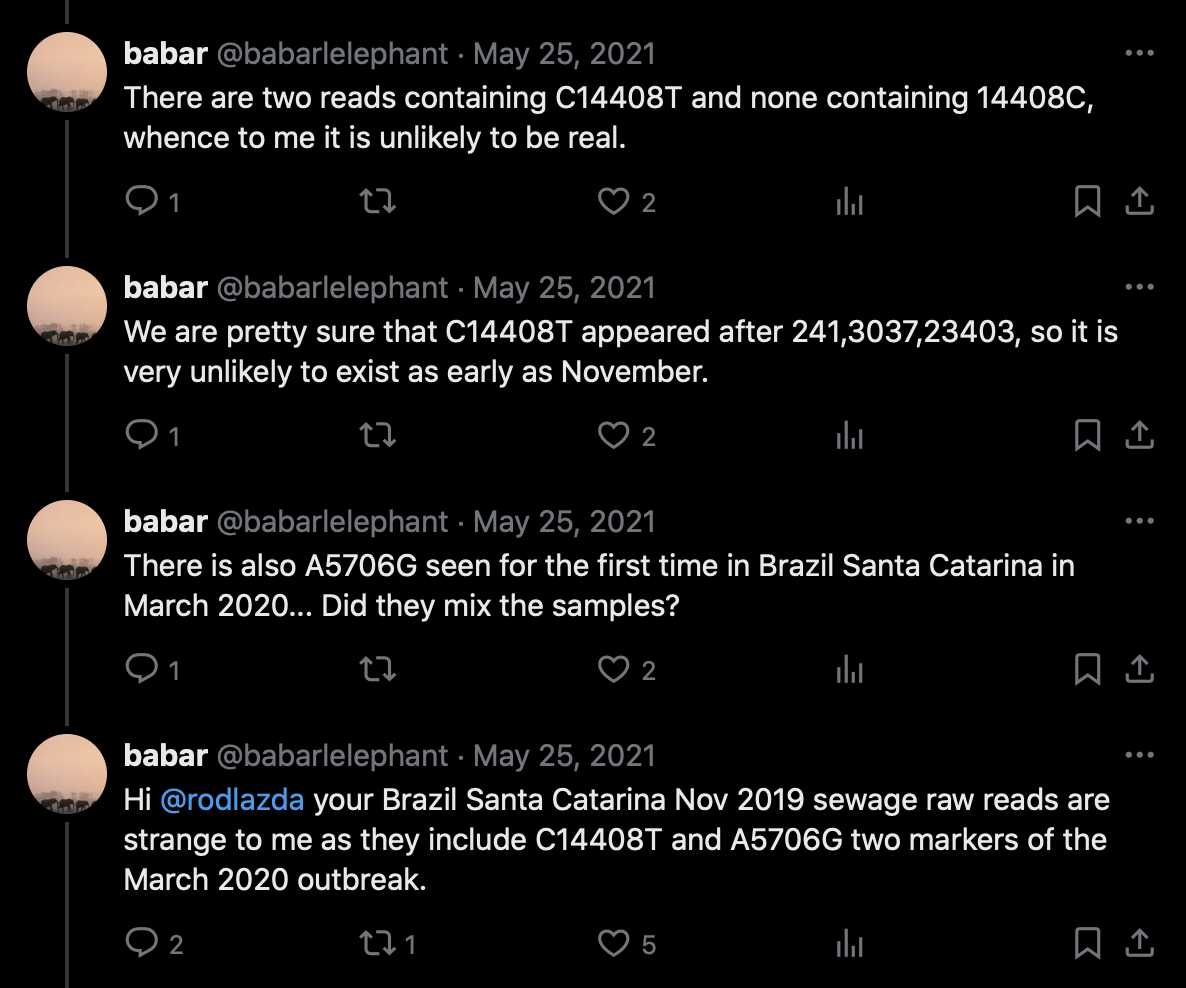
The abstract of a Brazilian paper from 2021 said: [https://www.sciencedirect.com/science/article/pii/S0048969721012651]
Human sewage from Florianopolis (Santa Catarina, Brazil) was analyzed for severe acute respiratory syndrome coronavirus-2 (SARS-CoV2) from October 2019 until March 2020. Twenty five ml of sewage samples were clarified and viruses concentrated using a glycine buffer method coupled with polyethylene glycol precipitation, and viral RNA extracted using a commercial kit. SARS-CoV-2 RNA was detected by RT-qPCR using oligonucleotides targeting N1, S and two RdRp regions. The results of all positive samples were further confirmed by a different RT-qPCR system in an independent laboratory. S and RdRp amplicons were sequenced to confirm identity with SARS-CoV-2. Genome sequencing was performed using two strategies; a sequence-independent single-primer amplification (SISPA) approach, and by direct metagenomics using Illumina's NGS. SARS-CoV-2 RNA was detected on 27th November 2019 (5.49 ± 0.02 log10 SARS-CoV-2 genome copies (GC) L−1), detection being confirmed by an independent laboratory and genome sequencing analysis. The samples in the subsequent three events were positive by all RT-qPCR assays; these positive results were also confirmed by an independent laboratory. The average load was 5.83 ± 0.12 log10 SARS-CoV-2 GC L−1, ranging from 5.49 ± 0.02 log10 GC L−1 (27th November 2019) to 6.68 ± 0.02 log10 GC L−1 (4th March 2020). Our findings demonstrate that SARS-CoV-2 was likely circulating undetected in the community in Brazil since November 2019, earlier than the first reported case in the Americas (21st January 2020).
Guy Gadboit downloaded the raw reads from the paper and wrote a Twitter thread about them. [https://x.com/gadboit/status/1793281441322693002]
Gadboit said that he only got 7% coverage for SARS2 even though the paper said that they got 25% coverage. However I got about 15% coverage:
$ enad()(printf %s\\n "${@-$(cat)}"|while IFS= read x;do curl -s "https://www.ebi.ac.uk/ena/portal/api/filereport?accession=$x&result=read_run&fields=fastq_ftp"|sed 1d|cut -f2|tr \; \\n;done|sed s,^,ftp://,|xargs wget -q)
$ x=SRR13152144;enad $x
$ fastp -35 -i $x\_1.fastq.gz -I $x\_2.fastq.gz -o $x\_1.fq.gz -O $x\_2.fq.gz
$ minimap2 -a --sam-hit-only sars2.fa $x\_[12].fq.gz|samtools sort ->$x.bam
$ samtools coverage $x.bam|column -t
#rname startpos endpos numreads covbases coverage meandepth meanbaseq meanmapq
MN908947.3 1 29903 92 4348 14.5403 0.293649 36.7 43.5
Gadboit said that he didn't see any other lineage-defining locations except D614G (A23403G) and C8782. However I actually didn't get any read that covered the position of D614G regardless of whether I trimmed reads or not. I got two reads for 8782 and two reads for 28144, but they both had the lineage B allele. But I didn't get any reads that covered the position of the third mutation of WA1 (18060) or the A0 mutation (29095):
$ samtools mpileup -f sars2.fa SRR13152144.bam|ruby -alne'next unless[8782,18060,28144,29095,23403].include?($F[1].to_i);next if$F[3]==0||$F[4]=="*";s=$F[4].upcase.gsub(/[.,]/,$F[2]).gsub(/\^./,"");s2=s.dup;s.enum_for(:scan,/[+-]\d+/).each{m=Regexp.last_match;m.begin(0).upto(m.end(0)+s[m.begin(0)+1,m.end(0)].to_i){|i|s2[i]=" "}};puts $F[1]+" "+$F[2]+" "+"ACGT".chars.map{|x|q=$F[5].chars.values_at(*s2.chars.each_index.select{|i|s2[i]==x}).map{|c|c.ord-33};q.size.to_s+(q.size==0?"":"["+"%.0f"%(q.sum*1.0/q.size)+"]")}*" "'|sed s/^/$x\ /|sort -n|(echo pos ref A C G T;cat)|column -t
pos ref A C G T
8782 C 0 2[34] 0 0
28144 T 0 0 0 2[38]
I got 4 different mutations with MAPQ over 10. The MAPQ of each mutation was around 50, because there were two high-quality reads that supported each mutation:
$ x=SRR13152144;samtools index $x.bam;bcftools mpileup -f sars2.fa $x.bam|bcftools call -mv|grep -v ^\#\#|cut -f-6|column -t #CHROM POS ID REF ALT QUAL MN908947.3 3392 . G C 3.22451 MN908947.3 5706 . A G 48.4146 MN908947.3 11437 . TAA TA 5.7555 MN908947.3 14408 . C T 48.4146 MN908947.3 14422 . C T 46.4146 MN908947.3 14895 . T G 47.4146
I don't understand why G3392C gets a MAPQ of only 3.2, even though
it's supported by two reads with high quality like the other mutations.
The quality of both bases is 39 (H, which is 72 minus
33):
$ samtools mpileup -f sars2.fa $x.bam|awk '$2~/^(3392|5706|14408|14422|14895)$/'|cut -f2-|column -t 3392 G 2 Cc HH 5706 A 2 Gg HH 14408 C 2 Tt HH 14422 C 2 Tt HF 14895 T 2 Gg HG
In a nearly complete set of GISAID submissions with a collection date in 2020, I didn't find any sample with all 4 mutations or even 3 out of 4 mutations, but the two earliest samples with 2 out of the 4 mutations were both from Santa Catarina in Brazil (which was the same state where the wastewater samples were collected):
$ curl -Ls sars2.net/f/gisaid2020.tsv.xz|xz -dc>gisaid2020.tsv
$ printf %s\\n A5706G C14408T C14422T T14895G|awk -F\\t 'NR==FNR{a[$0];next}{n=0;split($12,z,",");for(i in z)if(z[i] in a)n++;if(n>1)print n"\t"$0}' - gisaid2020.tsv|sort -k5|head|cut -f1-8,11-14|tr \\t \|
2|EPI_ISL_541370|hCoV-19/Brazil/SC-FIOCRUZ-770/2020|2020-09-22|2020-03-12|B.1.1.33|Brazil|Santa Catarina|Human|10|C241T,C3037T,A5706G,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C||
2|EPI_ISL_476278|hCoV-19/Brazil/SC-L16-CD314/2020|2020-06-25|2020-03-24|B.1.1.33|Brazil|Santa Catarina|Human|11|C241T,C3037T,A5706G,C14408T,G18803T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C||
2|EPI_ISL_470601|hCoV-19/Brazil/PA-0227/2020|2020-06-18|2020-04-01|B.1.1.33|Brazil|Para|Human|11|C241T,C3037T,A5706G,C14408T,A23403G,T27299C,A28119T,G28881A,G28882A,G28883C,T29148C||
2|EPI_ISL_2959070|hCoV-19/USA/MI-MDHHS-SC26889/2020|2021-07-15|2020-04-02|B.1|USA|Michigan|Human|9|C180T,C241T,C1059T,C3037T,C14408T,C14422T,A23403G,G25563T,G28314A||
2|EPI_ISL_447105|hCoV-19/USA/MI-MDHHS-SC20308/2020|2020-05-16|2020-04-02|B.1|USA|Michigan|Human|9|C180T,C241T,C1059T,C3037T,C14408T,C14422T,A23403G,G25563T,G28314A||
2|EPI_ISL_525869|hCoV-19/USA/OR-OHSU-0648/2020|2020-09-01|2020-06-16|B.1.134|USA|Oregon|Human|9|C241T,C3037T,A5706G,C8818T,A12579G,C14408T,C15952T,G20980A,A23403G||
2|EPI_ISL_525840|hCoV-19/USA/OR-OHSU-0546/2020|2020-09-01|2020-06-16|B.1.134|USA|Oregon|Human|9|C241T,C3037T,A5706G,C8818T,A12579G,C14408T,C15952T,G20980A,A23403G||
2|EPI_ISL_525870|hCoV-19/USA/OR-OHSU-0649/2020|2020-09-01|2020-06-16|B.1.134|USA|Oregon|Human|9|C241T,C3037T,A5706G,C8818T,A12579G,C14408T,C15952T,G20980A,A23403G||
2|EPI_ISL_576534|hCoV-19/USA/CA-IGI-0346/2020|2020-10-12|2020-06-29|B.1.1|USA|California|Human|11|C241T,C3037T,C14408T,C14422T,G21901A,A23403G,C24863T,C27145T,G28881A,G28882A,G28883C|686-694|
2|EPI_ISL_541116|hCoV-19/Spain/PV-IBV-98006728/2020|2020-09-22|2020-07-16|B.1|Spain|Basque Country|Human|13|C241T,C1392T,C1593T,T1623C,C3037T,C5512T,A5706G,G10870T,C12815T,C14408T,A20268G,A23403G,A27831G||
The Brazilian samples from Santa Catarina belonged to the clade B.1.1.33. They had D614G and 3 out of the 4 other B.1 mutations (with G25563T missing).
This shows samples whose set of mutation is a subset of the Brazilian sample with the earliest collection date, so that it shows different possible ancestor strains of the Brazilian sample. The first column shows how many times each mutation set was found in each country, and the second column shows how many total mutations the mutation set has relative to Wuhan-Hu-1. The samples with 9 shared mutations are mostly from Brazil, even though there's also samples from USA, Chile, Canada, Portugal, Australia, Uruguay, and UK. However 7 out of 10 mutations are shared by B.1.1 samples from many countries all over the world:
$ printf %s\\n C241T C3037T A5706G C14408T A23403G T27299C G28881A G28882A G28883C T29148C|awk -F\\t 'NR==FNR{a[$0];next}{split($12,b,",");for(i in b)if(!(b[i]in a))next;o[$11 FS$6 FS$5 FS$12]++}END{for(i in o)print o[i]FS i}' - gisaid2020.tsv|sort -rnk2|head -n50|column -ts$'\t'
1 10 Brazil B.1.1.33 C241T,C3037T,A5706G,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C
9 9 Chile B.1.1.33 C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C
7 9 Canada B.1.1.33 C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C
5 9 Portugal B.1.1.33 C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C
2 9 Australia B.1.1.33 C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C
159 9 Brazil B.1.1.33 C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C
11 9 USA B.1.1.33 C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C
1 9 Uruguay B.1.1.33 C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C
1 9 United Kingdom B.1.1.33 C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C
1 9 Brazil Unassigned C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C
6 8 Switzerland B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C,T29148C
4 8 Brazil B.1.1.33 C241T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C
3 8 Netherlands B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C,T29148C
3 8 Australia B.1.1.33 C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C
2 8 United Kingdom B.1.1.33 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C,T29148C
2 8 United Kingdom B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C,T29148C
2 8 Czech Republic B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C,T29148C
2 8 Brazil B.1.1 C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C
19 8 Brazil B.1.1.33 C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C
1 8 USA B.1.1.33 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C,T29148C
1 8 USA B.1.1 C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C
1 8 Italy B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C,T29148C
1 8 Brazil Unassigned C241T,C3037T,C14408T,T27299C,G28881A,G28882A,G28883C,T29148C
1 8 Brazil B.1.1.33 C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C
1 8 Brazil B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C,T29148C
1 8 Australia B.1.1.33 C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C
97 7 Russia B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
92 7 Italy B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
91 7 Spain B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
9 7 Turkey B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
9 7 Guam B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
9 7 Chile B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
8 7 United Arab Emirates B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
8 7 Poland B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
8 7 Finland B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
76 7 Germany B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
72 7 Australia B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
7 7 Madagascar B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
6 7 Peru B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
6 7 Mexico B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
6 7 Colombia B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
6 7 Bangladesh B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
56 7 Sweden B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
53 7 France B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
52 7 Canada B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
50 7 Cyprus B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
5 7 India B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
5 7 Hungary B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
5 7 Ecuador B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
4 7 South Korea B.1.1 C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
Out of the 5 SNVs I got when I ran mpileup on the November 2019 sample, only C14408T was part of the 7 mutations in B.1.1. The second mutation which matched the Brazilian samples on GISAID was A5706G, but the oldest two samples with the mutation are from the state of Santa Catarina in Brazil: [https://cov-spectrum.org/explore/World/AllSamples/from%3D2020-01-01%26to%3D2020-03-31/variants?nucMutations=A5706G]
$ grep A5706G gisaid2020.tsv|sort -t$'\t' -k4|head|cut -f1-8,11-14|tr \\t \| EPI_ISL_541370|hCoV-19/Brazil/SC-FIOCRUZ-770/2020|2020-09-22|2020-03-12|B.1.1.33|Brazil|Santa Catarina|Braco do Norte|10|C241T,C3037T,A5706G,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C|| EPI_ISL_476278|hCoV-19/Brazil/SC-L16-CD314/2020|2020-06-25|2020-03-24|B.1.1.33|Brazil|Santa Catarina||11|C241T,C3037T,A5706G,C14408T,G18803T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C|| EPI_ISL_470601|hCoV-19/Brazil/PA-0227/2020|2020-06-18|2020-04-01|B.1.1.33|Brazil|Para||11|C241T,C3037T,A5706G,C14408T,A23403G,T27299C,A28119T,G28881A,G28882A,G28883C,T29148C|| EPI_ISL_525869|hCoV-19/USA/OR-OHSU-0648/2020|2020-09-01|2020-06-16|B.1.134|USA|Oregon|Multnomah County|9|C241T,C3037T,A5706G,C8818T,A12579G,C14408T,C15952T,G20980A,A23403G|| EPI_ISL_525840|hCoV-19/USA/OR-OHSU-0546/2020|2020-09-01|2020-06-16|B.1.134|USA|Oregon|Washington County|9|C241T,C3037T,A5706G,C8818T,A12579G,C14408T,C15952T,G20980A,A23403G|| EPI_ISL_525870|hCoV-19/USA/OR-OHSU-0649/2020|2020-09-01|2020-06-16|B.1.134|USA|Oregon|Washington County|9|C241T,C3037T,A5706G,C8818T,A12579G,C14408T,C15952T,G20980A,A23403G|| EPI_ISL_541116|hCoV-19/Spain/PV-IBV-98006728/2020|2020-09-22|2020-07-16|B.1|Spain|Basque Country|San Sebastian|13|C241T,C1392T,C1593T,T1623C,C3037T,C5512T,A5706G,G10870T,C12815T,C14408T,A20268G,A23403G,A27831G|| EPI_ISL_13687819|hCoV-19/USA/WI-Mayo21940/2020|2022-07-06|2020-07-21|B.1.239|USA|Wisconsin||16|C241T,C3037T,A5706G,C7043T,C8917T,C10036T,C13945T,C14408T,G16245T,G16741A,G17334T,A19779G,A20268G,A23403G,G28378T,C28854T|| EPI_ISL_535485|hCoV-19/SouthAfrica/KRISP-K002622/2020|2020-09-15|2020-07-31|B.1.1.412|South Africa|KwaZulu-Natal||12|C241T,C3037T,C3165A,A5706G,G11447A,C14408T,C22675T,A23403G,G28077T,G28881A,G28882A,G28883C|| EPI_ISL_14291071|hCoV-19/Brazil/BA-FIOCRUZ-PVM6183/2020|2022-08-06|2020-08-03|B.1.1.33|Brazil|Bahia|Irece|15|C241T,C3037T,C3732T,A5706G,T13669G,C14408T,A23403G,T27299C,G27382C,A27383T,T27384C,G28881A,G28882A,G28883C,T29148C||
The Brazilian study also included a second SRA run, which was labeled
"Sewage metratranscriptome from Florianopolis, SC,
Brazil (Mar20)" (instead of "Nov19"
like the other run).
[https://trace.ncbi.nlm.nih.gov/Traces?run=SRR13152143]
bcftools call gave only 3 mutations for the March 2020
sample but none of them had a MAPQ above 10:
$ enad()(printf %s\\n "${@-$(cat)}"|while IFS= read x;do curl -s "https://www.ebi.ac.uk/ena/portal/api/filereport?accession=$x&result=read_run&fields=fastq_ftp"|sed 1d|cut -f2|tr \; \\n;done|sed s,^,ftp://,|xargs wget -q)
$ x=SRR13152143;enad $x
$ fastp -35 -i $x\_1.fastq.gz -I $x\_2.fastq.gz -o $x\_1.fq.gz -O $x\_2.fq.gz
$ minimap2 -a --sam-hit-only sars2.fa $x\_[12].fastq.gz|samtools sort ->$x.bam
$ samtools index $x.bam;bcftools mpileup -f sars2.fa $x.bam|bcftools call -mv|grep -v ^\#\#|cut -f2,4-6|column -t
POS REF ALT QUAL
6192 C T 3.7766
8145 A G 9.88514
8206 A G 9.88514
However all three mutations were rare in the set of GISAID submissions with a collection date in 2020, and none of them were found in samples from Brazil, and none of the submissions had even 2 out of the 3 mutations.
The March 2020 sample only got about 5% coverage for SARS2, and it didn't have any reads which covered the positions of lineage A (8782 and 28144), WA1 (18060), lineage A0 (29095), or D614G (23403):
$ samtools coverage SRR13152143.bam|column -t #rname startpos endpos numreads covbases coverage meandepth meanbaseq meanmapq MN908947.3 1 29903 23 1636 5.47102 0.0934689 37.4 57.8 $ samtools depth -a SRR13152143.bam|awk '$2~/^(8782|18060|28144|29095|23403)$/'|cut -f2- 8782 0 18060 0 23403 0 28144 0 29095 0
If I show all mutations instead of using bcftools call,
the March 2020 sample only gets one additional mutation (which is found
in 1 out of 2 reads and which has quality 18, so it's probably a
sequencing error):
$ x=SRR13152143;samtools mpileup -f sars2.fa $x.bam|puptab|awk '$2!=$3&&$4!=0'|(echo pos ref allele count coverage allele% average_quality;cat)|column -t pos ref allele count coverage allele% average_quality 6137 A C 1 2 50.000 18.0 6192 C T 1 1 100.000 31.0 8145 A G 1 1 100.000 39.0 8206 A G 1 1 100.000 39.0
In the set of GISAID submissions with a collection date in 2020, there's a total of 183 submissions where the state is listed as Santa Catarina. They include mutations in a total of 618 different positions, but none of hem are the 4 positions shown in the output above.
The March 2020 sample has zero depth for all positions where there's mutations in the two A5706G samples from Santa Catarina:
$ grep A5706G gisaid2020.tsv|grep Santa\ Catarina|cut -f12|tr , \\n|sed 's/.\(.*\)./\1/'|awk 'NR==FNR{a[$0];next}$2 in a' - <(samtools depth -a SRR13152143.bam)|cut -f2-|column -t
241 0
3037 0
5706 0
14408 0
18803 0
23403 0
27299 0
28881 0
28882 0
28883 0
29148 0
In the STAT results of the November 2019 sample, the most abundant leaf nodes were mostly human reads, human reads misclassified under monkeys, or bacteria:
$ curl "https://trace.ncbi.nlm.nih.gov/Traces/sra-db-be/run_taxonomy?acc=SRR13152144&cluster_name=public">SRR13152144.stat $ stal()(jq -r '[.[]|.tax_table[]|.parent]as$par|[.[]|.tax_table[]|select(.tax_id as$x|$par|index($x)|not)]|sort_by(-.total_count)[]|((.total_count|tostring)+";"+.org)' "$@") # STAT leaf nodes $ stal SRR13152144.stat|head -n24 29976;Homo sapiens 427;Escherichia coli 257;Gorilla gorilla gorilla 241;Klebsiella pneumoniae 185;Clostridioides difficile 171;Dolosigranulum pigrum (bacteria) 124;Pan troglodytes 115;Pongo abelii 97;unclassified Streptococcus 57;Salmonella enterica 53;Aotus nancymaae (monkey) 53;Pseudomonas aeruginosa (bacteria) 52;Bordetella bronchiseptica E013 (bacteria) 44;Nomascus leucogenys (monkey) 39;Streptococcus pneumoniae 35;Staphylococcus aureus 31;Haemophilus 26;Chlorocebus sabaeus (monkey) 20;Staphylococcus epidermidis (bacteria) 15;Veillonella sp. oral taxon 158 str. F0412 14;Severe acute respiratory syndrome coronavirus 2 11;Neisseria 8;Laurasiatheria 8;Oncorhynchus kisutch
After I had written the previous sections in this file, I found that babarlelephant had posted these tweets in 2021: [https://x.com/babarlelephant/status/1397204321838813199]

Half of the time when I think I have made some new discovery relating to sarbecoviruses, it has already been found long ago by Babar or Daoyu or breakfast_dogs.
The other 2 out of the 4 mutations with MAPQ above 10 don't seem to be very informative. The C14422T mutation seems to have emerged independently in a lineage A sample in China, in B.1 samples in USA, in D.2 samples in Australia, in B.1.237 samples in South Africa, and so on:
$ grep C14422T gisaid2020.tsv |awk -F\\t '!a[$12]++'|head|cut -f1,3-8,11-14|tr \\t \| EPI_ISL_412983|2020-03-02|2020-02-08|A|China|Hubei|Tianmen|8|A3175G,G3179A,C8782T,C14422T,C14585T,G23405C,T28144C,C28315T|| EPI_ISL_425457|2020-04-14|2020-04-01|B.29|United Kingdom|England||7|G11083T,C14422T,C14805T,C23707T,G26144T,G26779T,G29140C|| EPI_ISL_2959070|2021-07-15|2020-04-02|B.1|USA|Michigan||9|C180T,C241T,C1059T,C3037T,C14408T,C14422T,A23403G,G25563T,G28314A|| EPI_ISL_3085619|2021-07-26|2020-04-12|B.33|Sweden|Stockholm||8|G11083T,C14422T,C14805T,T17247C,C20428T,G26144T,G26849T,C29718A|| EPI_ISL_576534|2020-10-12|2020-06-29|B.1.1|USA|California|Contra Costa County|11|C241T,C3037T,C14408T,C14422T,G21901A,A23403G,C24863T,C27145T,G28881A,G28882A,G28883C|686-694| EPI_ISL_10560782|2022-03-02|2020-07-23|D.2|Australia|Victoria||22|C241T,A1163T,C3037T,A3193G,G7466T,T7540C,A13622G,C14408T,C14422T,G16647T,G17551T,C18555T,A21936G,G22992A,G23401A,A23403G,G28881A,G28882A,G28883C,C29187T,A29188G,C29870A|17564,24169,28899-28903|11095:T,13932:T,28148:T EPI_ISL_10560774|2022-03-02|2020-07-25|D.2|Australia|Victoria||20|C241T,G903T,A1163T,C3037T,T7540C,C14408T,C14422T,G15594T,G16647T,G17551T,C18555T,G21746T,A21936G,G22992A,G23401A,A23403G,C26625T,G28881A,G28882A,G28883C|17564,24169,28899-28903|13932:T,28148:T EPI_ISL_539717|2020-09-19|2020-08-19|B.1.1.306|India|Telangana||21|C241T,C313T,C3037T,G3871T,A4372G,C5700A,C9693T,C14408T,C14422T,C15342T,C16626T,A21550C,A21551T,C21646T,A23403G,C25460T,G26262T,G28326T,G28881A,G28882A,G28883C|| EPI_ISL_1040807|2021-02-21|2020-09-07|B.1.237|South Africa|Western Cape||18|G174T,C241T,C1205A,G2164C,C3037T,C6638T,C11455T,C12115T,C12439T,C14408T,C14422T,C15480T,G18651A,A20268G,A23403G,G23587C,C28854T,C29367T|| EPI_ISL_640120|2020-11-17|2020-09-15|B.1.237|South Africa|Western Cape|Western|21|G174T,C241T,C1205A,G2164C,C3037T,C6638T,C11455T,C12115T,C12439T,C14408T,C14422T,C15480T,T16483A,G16487C,G18651A,A20268G,A23403G,G23587C,C28760A,C28854T,C29367T||
There's 15 submissions with T14895G at covSPECTRUM. [https://cov-spectrum.org/explore/World/AllSamples/from%3D2020-01-01%26to%3D2024-12-31/variants?nucMutations=T14895G] In the set of GISAID submissions with a collection date in 2020, the T14895G mutation is found in only one sample that seems to have a lot of errors, because it has over 100 nucleotide changes and many long insertions:
$ grep T14895G gisaid2020.tsv|tr \\t \| EPI_ISL_935114|hCoV-19/Italy/CAM-CRGS-133/2020|2021-02-04|2020-11-13|B.1.177|Italy|Campania||30328|Human|118|T3C,A4C,G7T,C241T,T445C,C3037T,C6286T,C7851T,C14408T,T14459C,G14563A,C14564G,G14566A,G14569T,C14571T,C14576A,G14580C,G14584A,C14585T,C14588T,A14597G,T14598A,C14599T,T14603A,T14606G,A14612G,A14613C,C14614A,C14618T,C14625T,T14629A,C14630A,A14631T,G14632T,T14633C,G14635A,T14637A,C14639A,A14644C,G14656C,T14660A,T14661A,C14662G,T14667C,G14668C,C14670T,A14672G,C14675A,G14677A,T14679C,T14682A,T14683G,T14685C,A14687T,C14688A,A14689G,A14693T,C14697T,T14698G,A14699T,G14701A,C14703A,T14704A,G14707A,T14709C,T14711G,C14714T,A14716G,G14718C,G14719A,T14726G,T14727A,G14730A,A14732C,G14734C,A14736T,T14740C,G14746T,T14749C,A14752G,A14753C,C14755A,C14757A,T14762C,G14767A,G14773C,G14776C,T14778G,G14881T,A14882T,G14885T,T14887A,C14889T,G14893C,T14895G,C14898A,G14900A,T14904A,A14905C,T14907C,C14909A,A14912T,C14914T,G21255C,C22227T,G22346T,A23403G,C26801G,C28932T,G29645T,A29869C,C29870T,A29871G,A29874T,A29877G,A29878C,A29880T,A29882T|14590-14592,14647-14654|0:TTTTTTTTAATGATCGGGCGACCACCGACACATACACGAT,6:CACTCTTTCCCTACACGACGCGCTTCCGCTC,14457:AGA,14562:CTCCACCATCGTAACAATCAA,14572:AACAACTTCAACTACAAATAGTAGTTGT,14580:A,14607:TAGTAGTCATAATCGCT,14621:CATCCTGAGCAAAGAAGAA,14661:A,14723:GAAAAGCAACATTGTTAGTAAGTGCAGCTA,14739:GCGTTTATCTAGTAACACATAATCATCAC,14751:GCATTGTATGTTGAGA,14760:ATGAGG,14769:GTATGTGATGT,14774:CAAAATAATCACCAATATTTAATTTGTAAGTTGTTGTACC,14896:CGACCACC,14916:CACTATTTCCCTACACGACGCTCTTCCGATCT,19248:AGAGTGCTATCTAACCTTAACTTGCCTGGTTGTGATGGTGGCAGTTTGTATGTAAATAAACATGCATTCCACACACCAGCTTTTGATAAAAGTGCTTTTGTTAATTTAAAACAATTACCAAG,29866:TT|N:A220V,ORF1a:A2529V,ORF1b:P314L,ORF1b:F331R,S:A222V,S:A262S,S:D614G||20E|Europe|1
I thought that maybe the two mutations could've been sequencing errors if they were close to the ends of reads. But neither of them was close to the ends of reads. T14895G was close to the middle:
$ samtools view SRR13152144.bam|awk '$4+length($10)+20>=x&&$4<=x' x=14895|cut -f1,10|seqkit tab2fx|cat - sars2.fa|mafft --clustalout --globalpair -
[...]
SRR13152144.484 ---------------------------------------gactactatcgttataatcta
SRR13152144.484 ------------------------------------------------------------
MN908947.3 ttctttgctcaggatggtaatgctgctatcagcgattatgactactatcgttataatcta
SRR13152144.484 ccaacaatgtgtgatatcagacaactactatttgtagttgaagttgttgataagtacttt
SRR13152144.484 --------------------acaactactatttgtagttgaagttgttgataagtacttt
MN908947.3 ccaacaatgtgtgatatcagacaactactatttgtagttgaagttgttgataagtacttt
****************************************
SRR13152144.484 gattgttacgatgggggctgtattaatgctaaccaagtcatcgtcaacaacctagacaaa
SRR13152144.484 gattgttacgatgggggctgtattaatgctaaccaagtcatcgtcaacaacctagacaaa
MN908947.3 gattgttacgatggtggctgtattaatgctaaccaagtcatcgtcaacaacctagacaaa
************** *********************************************
SRR13152144.484 tcagctggt---------------------------------------------------
SRR13152144.484 tcagctggttttccatttaataaatggggtaaggctagactttattatga----------
MN908947.3 tcagctggttttccatttaataaatggggtaaggctagactttattatgattcaatgagt
*********
[...]
Zach Hensel whose wrote: "I see that one T->C is T16251C which, like A5706G, was also found in Brazil in a B.1.1.33 lineage sample in 2020." [https://x.com/alchemytoday/status/1793313402657308946]
However T16251C is only supported by 1 out of 2 reads that cover the position and it has low quality (16):
$ samtools view SRR13152144.bam|awk '$4+length($10)+20>=x&&$4<=x' x=16251|cut -f1,10|seqkit tab2fx|
cat - sars2.fa|mafft --clustalout --globalpair -
[...]
SRR13152144.194 -----------------------------------------------------------t
SRR13152144.194 -----------------------------------------------------------t
MN908947.3 gacatgtattctgttatgcttactaatgataacacttcaaggtattgggaacctgagttt
*
SRR13152144.194 tatgaagctatgtacacaccgcatacagtcttacaggctgttggggcttgcgttctttgc
SRR13152144.194 tatgaggctatgtacacaccgcatacagacttacaggctgttggggcttgtgttctttgc
MN908947.3 tatgaggctatgtacacaccgcatacagtcttacaggctgttggggcttgtgttctttgc
*****.********************** *********************.*********
^ T16251C
SRR13152144.194 aattcacagacttcattaaga---------------------------------------
SRR13152144.194 aattcacagacttcattaaga---------------------------------------
MN908947.3 aattcacagacttcattaagatgtggtgcttgcatacgtagaccattcttatgttgtaaa
*********************
[...]
On GISAID there's only two samples from Brazil which have T16251C. [https://cov-spectrum.org/explore/Brazil/AllSamples/AllTimes/variants?nucMutations=T16251C] One sample is from 2021. The other is B.1.1.33 like the samples with A5706G, but its collection date is only in July 2020. A5706G is found in two sets of B.1.1.33 samples from Brazil: there's 3 samples with collection dates in March-April 2020 from Santa Catarina and Para, and there's also 7 samples with a collection date in August to September 2020 from Bahia. However neither set has T16251C.
The November 2019 run has a total of 25 positions with a mutation relative to Wuhan-Hu-1 if I also include mutations that are only supported by a single low-quality read. Out of 20 mutations that are only supported by a single read, all except 2 have a quality of 20 or below:
$ puptab()(ruby -ane's=$F[4].upcase.gsub(/[.,]/,$F[2]).gsub(/\^./,"").gsub("$","");s2=s.dup;s.enum_for(:scan,/[+-]\d+/).each{m=Regexp.last_match;m.begin(0).upto(m.end(0)+s[m.begin(0)+1,m.end(0)].to_i-1){|i|s2[i]=" "}};s2.gsub!(" ","");t=s2.chars.tally;"ACGT".chars.map{|x|q=$F[5].chars.values_at(*s2.chars.each_index.select{|i|s2[i]==x}).map{|c|c.ord-33};puts [$F[1],$F[2],x,t[x]||0,s2.size,"%.3f"%(100.0*(t[x]||0)/s2.size),q.size==0?"NA":"%.1f"%(q.sum.to_f/q.size)]*" "}' "$@")
$ x=SRR13152144;samtools mpileup -f sars2.fa $x.bam|puptab|awk '$2!=$3&&$4!=0'|(echo pos ref allele count coverage allele% average_quality;cat)|column -t
pos ref allele count coverage allele% average_quality
3392 G C 2 2 100.000 39.0
5706 A G 2 2 100.000 39.0
5929 T G 1 1 100.000 24.0
8576 A C 1 2 50.000 16.0
8601 T C 1 2 50.000 16.0
8755 A T 1 2 50.000 18.0
14408 C T 2 2 100.000 39.0
14422 C T 2 2 100.000 38.0
14895 T G 2 2 100.000 38.5
15106 A C 1 2 50.000 14.0
15125 T C 1 2 50.000 18.0
15132 T A 1 2 50.000 20.0
15133 A C 1 2 50.000 18.0
15168 G A 1 2 50.000 16.0
15172 T C 1 2 50.000 18.0
15189 A C 1 2 50.000 17.0
15195 T G 1 2 50.000 14.0
16206 G A 1 2 50.000 20.0
16229 T A 1 2 50.000 17.0
16251 T C 1 2 50.000 16.0
17890 A G 1 4 25.000 18.0
17893 G A 1 3 33.333 13.0
19218 T G 1 1 100.000 15.0
19273 C G 1 1 100.000 16.0
20488 T G 1 3 33.333 37.0
However the low-quality mutations listed above didn't include any more mutations which were found in Brazilian B.1.1.33 samples:
$ x=SRR13152144;samtools mpileup -f sars2.fa $x.bam|puptab|awk '$2!=$3&&$4!=0'|awk 'NR==FNR{a[$0];next}$2$1$3 in a' <(grep B\.1\.1\.33 gisaid2020.tsv|grep Brazil|cut -f12|tr , \\n) -|(echo pos ref allele count coverage allele% average_quality;cat)|column -t
pos ref allele count coverage allele% average_quality
5706 A G 2 2 100.000 39.0
14408 C T 2 2 100.000 39.0
16251 T C 1 2 50.000 16.0
Positions where B.1.1.33 has mutations had zero depth apart from 14408:
$ awk '$5=="B.1.1.33"' gisaid2020.tsv|cut -f12|awk '{split($0,z,",");for(i in z)a[z[i]]++}END{for(i in a)if(a[i]>NR/2)print i}'|sed 's/.\(.*\)./\1/'|awk 'NR==FNR{a[$0];next}$2 in a' - <(samtools depth -a SRR13152144.bam)|cut -f2-|column -t
241 0
3037 0
14408 2
23403 0
27299 0
28881 0
28882 0
28883 0
29148 0
Apart from 5706 and 14408, there was also zero depth for all positions with mutations in the A5706G samples from Santa Catarina:
$ grep A5706G gisaid2020.tsv|grep Santa\ Catarina|cut -f12|tr , \\n|sed 's/.\(.*\)./\1/'|awk 'NR==FNR{a[$0];next}$2 in a' - <(samtools depth -a SRR13152144.bam)|cut -f2-|column -t
241 0
3037 0
5706 2
14408 2
18803 0
23403 0
27299 0
28881 0
28882 0
28883 0
29148 0
Gadboit posted this tweet: [https://x.com/gadboit/status/1794000682669998429]
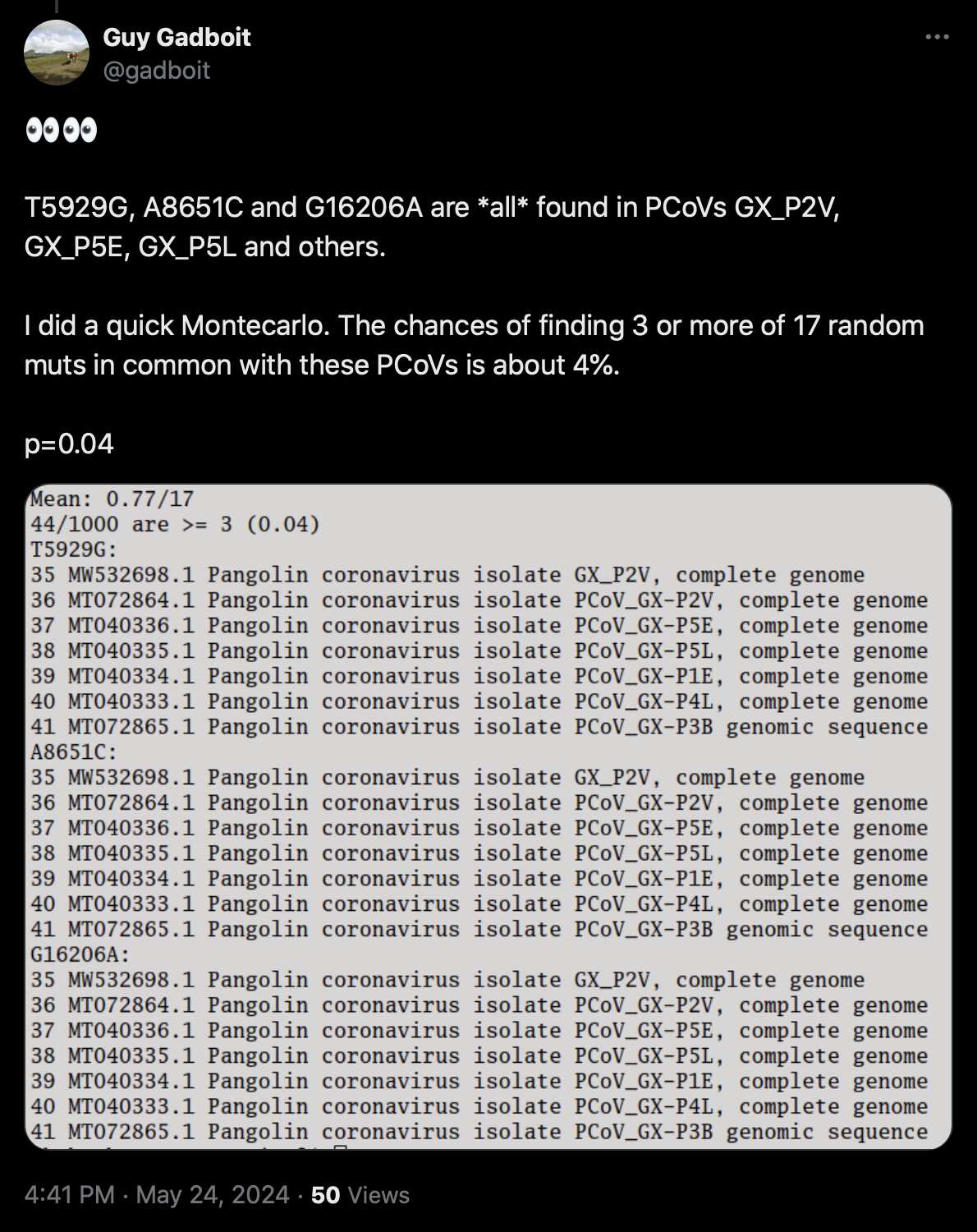
A8561C is probably a sequencing error, because it's on the sixth-last
base of the read and the base quality is 17. That part of the read got
trimmed when I used fastp -35, because it removed segments
with low-quality bases from the ends of reads. So I got zero coverage
for 8651 when I used trimmed reads:
$ puptab()(ruby -ane's=$F[4].upcase.gsub(/[.,]/,$F[2]).gsub(/\^./,"").gsub("$","");s2=s.dup;s.enum_for(:scan,/[+-]\d+/).each{m=Regexp.last_match;m.begin(0).upto(m.end(0)+s[m.begin(0)+1,m.end(0)].to_i-1){|i|s2[i]=" "}};s2.gsub!(" ","");t=s2.chars.tally;"ACGT".chars.map{|x|q=$F[5].chars.values_at(*s2.chars.each_index.select{|i|s2[i]==x}).map{|c|c.ord-33};puts [$F[1],$F[2],x,t[x]||0,s2.size,"%.3f"%(100.0*(t[x]||0)/s2.size),q.size==0?"NA":"%.1f"%(q.sum.to_f/q.size)]*" "}' "$@")
$ samtools mpileup -f sars2.fa SRR13152144.bam|puptab|awk '$1~/^(5929|8651|16206)$/&&$4'|(echo pos ref al count depth al% average_quality;cat)|column -t
pos ref al count depth al% average_quality
5929 T G 1 1 100.000 24.0
16206 G A 1 2 50.000 20.0
16206 G G 1 2 50.000 16.0
When I didn't trim the reads, I got one read that had A8651C, but the quality of the base with the mutation was 17:
$ minimap2 -a --sam-hit-only sars2.fa SRR13152144_[12].fastq.gz|samtools sort ->SRR13152144.untrim.bam [...] $ samtools mpileup -f sars2.fa SRR13152144.untrim.bam|puptab|awk '$1~/^(5929|8651|16206)$/&&$4'|(echo pos ref al count depth al% average_quality;cat)|column -t pos ref al count depth al% average_quality 5929 T G 1 1 100.000 24.0 8651 A C 1 1 100.000 17.0 16206 G A 1 2 50.000 20.0 16206 G G 1 2 50.000 16.0
The output above shows that G16206A is supported by only one read out of two reads that cover the position, and the quality of the base with the mutation is 20. And the 5929 position is covered by only one read, and the quality of the base with the mutation at 5929 is 24.
So all three of Gadboit's pangolin mutations are likely to be sequencing errors.
This shows that the base with a mutation at 8651 is also surrounded
by other low-quality bases, so it got removed by fastp -35
which removes segments with low-quality bases from the ends of
reads:
$ samtools view SRR13152144.untrim.bam|awk '$4>8500&&$4<=8651'|cut -f-12|tr \\t \|
SRR13152144.253277|0|MN908947.3|8564|30|93M57S|*|0|0|AATAATTGGTTGCAGCAGTTAATTAAAGTTACACTTGCGTTCCTTTTTGTTGCTGCTATTTTCTATTTAATAACACCTGTTCATGTCCTGTCTCTTATACTCTTCTGACGCTGCCGACGATTTGCAGTATTTTGAACTCGGATGATGCCT|>>A113B3BFAA11BBA113B3FFG1313D3FGFBGG1AAEFBGDGGDAGGA11B1B1DDHEBGEFHH22D22BFBEGFAFF222D22FFB1@2FF1@E22@2B1@12B//BEEF/>/>/F/F122111>2F22/11B1/////1B2BB1|NM:i:3
SRR13152144.253277|16|MN908947.3|8564|60|63S87M|*|0|0|GAAGAAGACGGAATACGAGATGTAGAGAGGTATCGTGGGCTCGGAGATGTGTATAAGAGACAGAATAATTGGTTGAAGCAGTTAATTAAAGTTACACTTGTGTTCCTTTTTGTTGCTGCTATTTTCTATTTAATAACACCTGTTCATGTC|>122<B0102B1B/?0>>211FHFCF11B///>/F/?//>>>1@G1GEG2B2F1HG2HHHFHHHHHGEGCB1D1HGB11HFEF2GFADHHGGHFFD0FCFHFB000B000GGCGHHHGFEF0A1A1F3BGGGGGA1BFAFFFFDFA>11A|NM:i:0
$ samtools view SRR13152144.untrim.bam|awk '$4>8500&&$4<=8651'|head -n1|cut -f10,11|tr \\t \\n|cut -c-93
AATAATTGGTTGCAGCAGTTAATTAAAGTTACACTTGCGTTCCTTTTTGTTGCTGCTATTTTCTATTTAATAACACCTGTTCATGTCCTGTCT
>>A113B3BFAA11BBA113B3FFG1313D3FGFBGG1AAEFBGDGGDAGGA11B1B1DDHEBGEFHH22D22BFBEGFAFF222D22FFB1@
the base at position 8651 has quality 2 = 50-33 = 17 ^
other surrounding bases also have low quality ^^^ ^^ ^
Gadboit also posted this tweet: [https://x.com/gadboit/status/1794284842060034514]
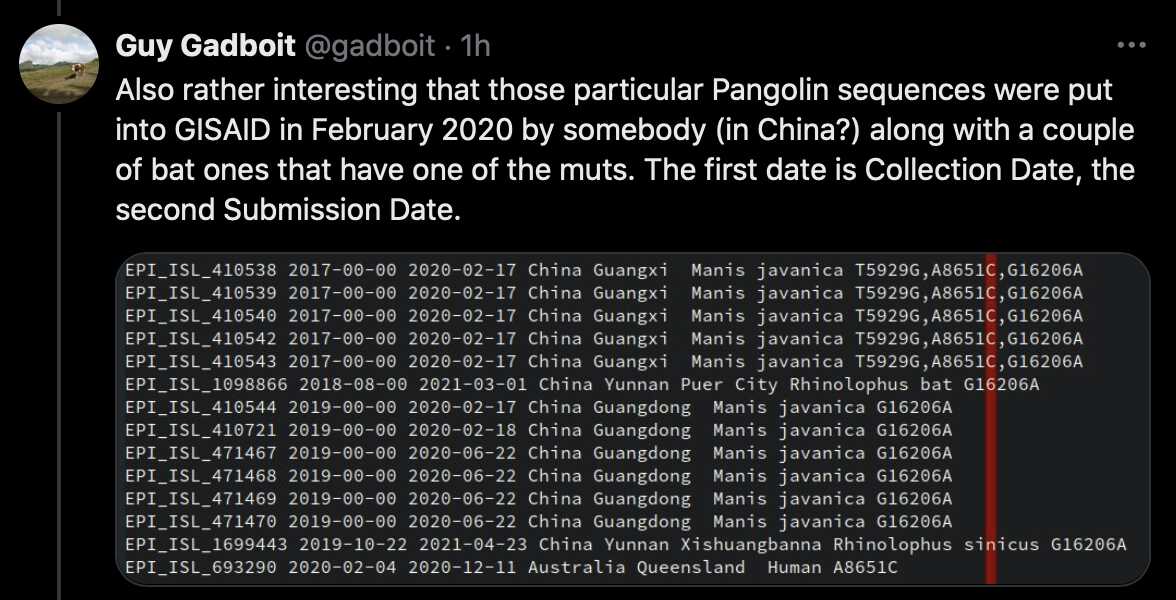
I didn't know if he meant that the bat sequences in his screenshot were also submitted in February 2020, because actually their submission date was in 2021. But Gadboit answered that he only meant that the pangolin sequences were submitted in February 2020.
I told him that the first two mutations are also in Rc-os20. And mutations 1 and 3 are in GZ2021C, GZ2021H, and GZ2021I. And mutations 2 and 3 are in Rc-mk2 and in 10 different Vietnamese sequences from Hassanin et al. 2023:
brew install --cask miniconda;conda init ${SHELL##*/}
conda install -c bioconda nextclade
nextclade dataset get --name sars-cov-2 --output-dir sars-cov-2
curl -sL sars2.net/f/sarbe.fa.xz|gzip -dc>sarbe.fa
nextclade run --input-dataset sars-cov-2 --output-all cladeout <(seqkit seq -g sarbe.fa)
awk -F\\t '$26~/T5929G/&&$26~/A8651C/' cladeout/nextclade.tsv|cut -f2
Gadboit answered: "Yes I've also found some of the muts in those Japanese bat viruses Rc-mk2 and Rc-os20. There are actually four of the 17 in each, but a different four. The PCoVs still seem to be the best match." [https://x.com/gadboit/status/1794309583063978444]
My maximum number of shared mutations was only 3/25, which might be because I trimmed the reads, but there were 15 different sequences on shared first place. Here the first column shows the number of mutations shared with the Brazilian reads, and the second column shows total nucleotide changes from Wuhan-Hu-1 with indels not included:
$ samtools mpileup -f sars2.fa SRR13152144.bam|puptab|awk '$2!=$3&&$4{print$2$1$3}'|awk -F\\t 'NR==FNR{a[$0];next}{n=0;for(i in a)if($26~i)n++;print n,$11,$2}' - <(sed 1d cladeout/nextclade.tsv)|sort -rn|sed -E 's/( orf1ab| surface|,).*//'|head -n20
3 6554 OP837780.1 UNVERIFIED: Sarbecovirus sp. isolate RhGB03
3 5957 DQ412042.1 Bat SARS coronavirus Rf1
3 5893 MK211374.1 Coronavirus BtRl-BetaCoV/SC2018
3 5829 GQ153542.1 Bat SARS coronavirus HKU3-7
3 5815 OK017811.1 Sarbecovirus sp. isolate FJ2021M
3 5805 OK017810.1 Sarbecovirus sp. isolate FJ2021E
3 5805 OK017808.1 Sarbecovirus sp. isolate FJ2021A
3 5794 GQ153543.1 Bat SARS coronavirus HKU3-8
3 5781 OK017809.1 Sarbecovirus sp. isolate FJ2021D
3 5778 OK017807.1 Sarbecovirus sp. isolate AH2021A
3 5528 KF294457.1 Bat SARS-like coronavirus isolate Longquan-140
3 5481 LC663958.1 Sarbecovirus sp. Rc-os20 RNA
3 5417 LC663959.1 Sarbecovirus sp. Rc-mk2 RNA
3 3481 MG772934.1 Bat SARS-like coronavirus isolate bat-SL-CoVZXC21
3 3353 OQ297732.1 MAG: Severe acute respiratory syndrome-related coronavirus isolate GD/L105.18/2022
2 5999 KY770860.1 Sarbecovirus sp. isolate Jiyuan-84
2 5932 DQ648856.1 Bat coronavirus (BtCoV/273/2005)
2 5929 KJ473812.1 BtRf-BetaCoV/HeB2013
2 5921 KJ473813.1 BtRf-BetaCoV/SX2013
2 5910 MZ081377.1 Betacoronavirus sp. RmYN07 strain bat/Yunnan/RmYN07/2020
The November 2019 reads cover 3 out of 12 positions where B.1.1.33 has a mutation relative to WA1/proCoV2, but all 3 positions have the allele in B.1.1.33 (which is also the allele in B.1), and all 3 positions are covered by two reads with a high base quality:
$ curl -Ls sars2.net/f/gisaid2020.tsv.xz|xz -dc>gisaid2020.tsv
$ gicon()(cut -f12 "$@"|awk '{split($0,z,",");for(i in z)a[z[i]]++}END{for(i in a)if(a[i]>NR/2)print i}')
$ awk -F\\t '$5=="B.1.1.33"' gisaid2020.tsv|gicon # nucleotide changes in more than 50% of B.1.1.33 samples
G28881A
G28882A
G28883C
C14408T
C3037T
C241T
T29148C
T27299C
A23403G
$ samtools mpileup -f sars2.fa SRR13152144.bam|awk 'NR==FNR{a[$0];next}$2 in a' <(awk -F\\t '$5=="B.1.1.33"' gisaid2020.tsv|gicon|sed 's/.\(.*\)./\1/';printf %s\\n 8782 18060 28144) -|column -t
MN908947.3 8782 C 2 ., BE
MN908947.3 14408 C 2 Tt HH
MN908947.3 28144 T 2 ., HG
Gadboit posted this table which shows all positions with mutations in WA1 or the B.1.1.33 consensus: [https://x.com/gadboit/status/1795723769878843738]
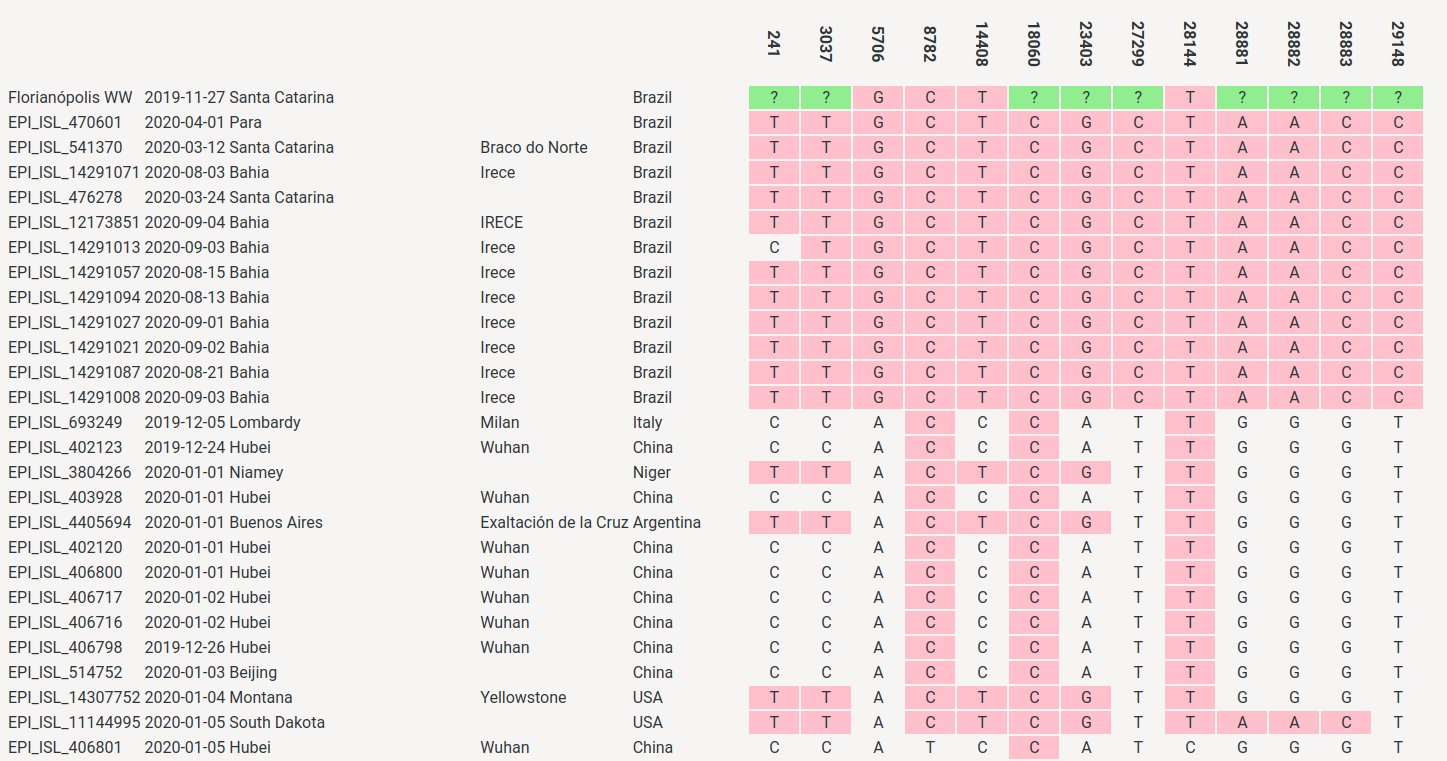
I made a similar table, but I also added the other 3/5 mutations that were called in the November 2019 sample, and I added mutations that were not called in the November 2019 but that were supported by 1/1 reads, and I added the 3 mutations that were called in the March 2020 sample, and I added other mutations that were found in early B.1.1.33 samples from Santa Catarina on GISAID:

In the set of GISAID submissions with a collection date in 2020, B.1.1.33 is actually by far the most common in Brazil:
$ awk -F\\t '$5=="B.1.1.33"{a[$6]++}END{for(i in a)print a[i],i}' gisaid2020.tsv|sort -rn|head
3366 Brazil
162 USA
71 Chile
51 Uruguay
32 Canada
22 United Kingdom
18 Argentina
16 Paraguay
16 Australia
12 French Guiana
The oldest B.1.1.33 sample by collection date is also from Brazil (even though it might have an incorrect collection date):
$ awk -F\\t '$5=="B.1.1.33"{a[$4" "$6]++}END{for(i in a)print a[i],i}' gisaid2020.tsv|sort -k2|head
1 2020-03-01 Brazil
1 2020-03-07 USA
1 2020-03-07 United Kingdom
1 2020-03-09 Brazil
1 2020-03-09 United Kingdom
1 2020-03-11 Brazil
3 2020-03-12 Brazil
1 2020-03-12 United Kingdom
4 2020-03-13 Brazil
1 2020-03-13 Canada
There's only 5 B.1.1.33 submissions that have an older or same collection date than the earliest submission with A5706G, but 3 out of 5 of them are from Brazil. There's 289 B.1.1 samples with an older or same collection date but only 2 of them are from Brazil. B.1.1.33 is defined by the mutations T27299C and T29148C, where the second mutation might to have arisen earlier, because in the output below the samples that are intermediate to B.1.1 and B.1.1.33 only have the second mutation but not the first one, and the intermediate samples are only found in Europe:
$ reline()(gawk -F\\t '{a[$12][$5]++}END{for(i in a){max=0;for(j in a[i])if(a[i][j]>max){max=a[i][j];out=j FS i}print out}}' ${1-~/gisaid2020.tsv}) # show most common lineage designation for each mutation set, or the first listed designation for ties
$ gian()(cut -f12|tr , \\n|awk -F\\t 'NR==FNR{a[$0];next}{split($12,b,",");for(i in b)if(!(b[i]in a))next}1' - ${1-~/gisaid2020.tsv}) # keep only possible ancestor sequences whose mutation set is a subset of the list of mutations from STDIN
$ grep A5706G gisaid2020.tsv|head -n1|gian|awk -F\\t '$11>=5&&$4<="2020-03-12"'|awk -F\\t 'NR==FNR{a[$1]=$2;next}{b[$11 FS a[$12]FS$6 FS$12]++}END{for(i in b)print b[i]FS i}' <(reline) -|sort -rnk2|csvtk -t pretty -s\ |sed 2d
1 10 B.1.1.33 Brazil C241T,C3037T,A5706G,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C
2 9 B.1.1.33 Brazil C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C
1 9 B.1.1.33 United Kingdom C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C
1 9 B.1.1.33 USA C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C
5 8 B.1.1 Switzerland C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C,T29148C
3 8 B.1.1 Netherlands C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C,T29148C
2 8 B.1.1 United Kingdom C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C,T29148C
1 8 B.1.1 Italy C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C,T29148C
1 8 B.1.1 Czech Republic C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C,T29148C
9 7 B.1.1 Vietnam C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
9 7 B.1.1 Denmark C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
86 7 B.1.1 United Kingdom C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
8 7 B.1.1 Switzerland C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
6 7 B.1.1 Netherlands C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
6 7 B.1.1 Israel C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
5 7 B.1.1 Spain C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
5 7 B.1.1 Portugal C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
5 7 B.1.1 Japan C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
41 7 B.1.1 Italy C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
4 7 B.1.1 Germany C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
3 7 B.1.1 New Zealand C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
2 7 B.1.1 Mexico C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
2 7 B.1.1 Ireland C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
2 7 B.1.1 France C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
2 7 B.1.1 Brazil C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
2 7 B.1.1 Austria C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
19 7 B.1.1 USA C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
16 7 B.1.1 Sweden C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
15 7 B.1.1 Greece C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
12 7 B.1.1 Belgium C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
1 7 B.1.1 Sri Lanka C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
1 7 B.1.1 Singapore C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
1 7 B.1.1 Saudi Arabia C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
1 7 B.1.1 Morocco C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
1 7 B.1.1 Lebanon C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
1 7 B.1.1 Cyprus C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
1 7 B.1.1 Canada C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
1 7 B.1.1 Bolivia C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
1 7 B.1.1 Australia C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
4 6 B.1.1 Australia C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
1 6 B.1.1 United Kingdom C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
1 6 B.1.1 Russia C241T,C3037T,A23403G,G28881A,G28882A,G28883C
1 6 B.1.1 Ireland C3037T,C14408T,A23403G,G28881A,G28882A,G28883C
1 6 B.1.1 Ireland C241T,C14408T,A23403G,G28881A,G28882A,G28883C
1 5 B.1.1 Ireland C3037T,A23403G,G28881A,G28882A,G28883C
However when I didn't exclude samples whose collection date was later than the earliest submission with A5706G, then there were also intermediate samples from Brazil that only had the first mutation but not the second one:
$ grep A5706G gisaid2020.tsv|head -n1|gian|awk -F\\t '$11>=5'|awk -F\\t 'NR==FNR{a[$1]=$2;next}{b[$11 FS a[$12]FS$6 FS$12]++}END{for(i in b)print b[i]FS i}' <(reline) -|awk '$2==8'|csvtk -t pretty -s\ |sed 2d
1 8 USA C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C
1 8 Italy C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C,T29148C
3 8 Netherlands C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C,T29148C
4 8 United Kingdom C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C,T29148C
2 8 Czech Republic C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C,T29148C
4 8 Brazil C241T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C
3 8 Australia C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C
6 8 Switzerland C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C,T29148C
1 8 Brazil C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C,T29148C
19 8 Brazil C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C
1 8 Australia C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C
1 8 Brazil C241T,C3037T,C14408T,T27299C,G28881A,G28882A,G28883C,T29148C
3 8 Brazil C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C
1 8 USA C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C,T29148C
The oldest B.1.1.33 submission from Brazil is also unusual because it has a distance of 6 to its closest neighbors on GISAID, because it has 6 mutations that are missing from other B.1.1.33 samples (except there's a couple of Brazilian B.1.1.33 samples that contain 1 out of 6 of the mutations):
$ awk -F\\t '$5=="B.1.1.33"' gisaid2020.tsv|sed 1q|gicon|gidi|awk '$1<8'|sort -n|head|cut -f1,2,5-8,12,13|tr \\t \||pc 0|EPI_ISL_1739298|2020-03-01|B.1.1.33|Brazil|Rio de Janeiro|15|C241T,C3037T,C5183T,C6113T,C8668T,C14408T,G18756T,T21066A,A23403G,C23683T,T27299C,G28881A,G28882A,G28883C,T29148C 6|EPI_ISL_10553069|2020-04-06|B.1.1.33|Canada|Ontario|9|C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C 6|EPI_ISL_10553070|2020-04-03|B.1.1.33|Canada|Ontario|9|C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C 6|EPI_ISL_1068321|2020-04-25|B.1.1.33|Brazil|Bahia|9|C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C 6|EPI_ISL_1068391|2020-04-23|B.1.1.33|Brazil|Bahia|9|C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C 6|EPI_ISL_1139068|2020-03-25|B.1.1.33|Brazil|Mato Grosso do Sul|9|C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C 6|EPI_ISL_1181356|2020-04-25|B.1.1.33|Brazil|Ceara|9|C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C 6|EPI_ISL_1181594|2020-04-20|B.1.1.33|Brazil|Rio de Janeiro|9|C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C 6|EPI_ISL_1181603|2020-04-13|B.1.1.33|Brazil|Rio de Janeiro|9|C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C 6|EPI_ISL_1181606|2020-04-05|B.1.1.33|Brazil|Rio de Janeiro|9|C241T,C3037T,C14408T,A23403G,T27299C,G28881A,G28882A,G28883C,T29148C
However the 6 mutations aren't even that rare, because one of them is found in only 5 submissions, but the others are all found in at least a hundred submissions:
$ awk -F\\t '$5=="B.1.1.33"' gisaid2020.tsv|sed 1q|cut -f12|tr , \\n|sort|comm -23 - <(awk -F\\t '$5=="B.1.1.33"' gisaid2020.tsv|gicon|sort)|awk 'NR==FNR{a[$0];next}$0 in a' - <(cut -f12 gisaid2020.tsv|tr , \\n)|sort|uniq -c|sort -n
5 T21066A
163 C6113T
182 C8668T
886 G18756T
1321 C23683T
1662 C5183T
There's one Brazilian submission from July 2020 and another submission from September 2020 which both share 3 out of 6 of the mutations, and there's also a bunch of pangolin viruses which share 3 out of 6 of the mutations (even though 2 mutations are also shared by several RmYN and RsYN bat viruses which are not shown here):
$ gisum()(awk -F\\t 'NR==FNR{a[$0];next}{split($12,z,",");n=0;for(i in z)if(z[i]in a)n++;print n FS$0}' - ${1-~/gisaid2020.tsv})
$ printf %s\\n T21066A C6113T C8668T G18756T C23683T C5183T|gisum|awk '$1>1'|sort -rn|cut -f-12|head|tr \\t \|
6|EPI_ISL_1739298|hCoV-19/Brazil/RJ-IC0102020_158-20/2020|2021-04-26|2020-03-01|B.1.1.33|Brazil|Rio de Janeiro||29796|Human|15
3|EPI_ISL_5416113|hCoV-19/Brazil/MG-LBIb2964/2020|2021-10-21|2020-07-15|N.4|Brazil|Minas Gerais|Betim|29393|Human|20
3|EPI_ISL_410543|hCoV-19/pangolin/Guangxi/P3B/2017|2020-02-17|2017|B.1|China|Guangxi||29801|Manis javanica|3888
3|EPI_ISL_410542|hCoV-19/pangolin/Guangxi/P2V/2017|2020-02-17|2017|B.1|China|Guangxi||29795|Manis javanica|4249
3|EPI_ISL_410540|hCoV-19/pangolin/Guangxi/P5L/2017|2020-02-17|2017|B.1|China|Guangxi||29806|Manis javanica|4253
3|EPI_ISL_410539|hCoV-19/pangolin/Guangxi/P1E/2017|2020-02-17|2017|B.1|China|Guangxi||29801|Manis javanica|4257
3|EPI_ISL_410538|hCoV-19/pangolin/Guangxi/P4L/2017|2020-02-17|2017|B.1|China|Guangxi||29806|Manis javanica|4255
3|EPI_ISL_2557352|hCoV-19/Brazil/MG-FIOCRUZ-34167/2020|2021-06-16|2020-09-01|B.1.1.33|Brazil|Minas Gerais|Itaobim|29719|Human|20
2|EPI_ISL_848343|hCoV-19/USA/IL-IDPH-I-000137/2020|2021-01-19|2020-10-08|B.1.578|USA|Illinois|Cook County|29782|Human|22
2|EPI_ISL_848343|hCoV-19/USA/IL-IDPH-I-000137/2020|2021-01-19|2020-10-08|B.1.578|USA|Illinois|Cook County|29782|Human|22
B.1.1 is defined by three consecutive nucleotide changes at positions 28881, 28882, and 28883. If samples that likely have an incorrect collection date are excluded, the earliest sample by collection date with all three mutations is from Moscow, the next sample is from China, and the next 3 samples are from UK, so it's difficult to tell where B.1.1 may have originated. The samples from China or Moscow might also have an incorrect collection date. Most of the earliest sequences are from Europe, but there's also two sequences from Brazil with a collection date on February 26th:
$ grep G28881A,G28882A,G28883C gisaid2020.tsv|awk '$5=="B.1.1"&&!$NF'|head|cut -f1,3-8,11,12|tr \\t \| EPI_ISL_8907761|2022-01-20|2020-02-03|B.1.1|Russia|Moscow|Moscow|6|C241T,C3037T,A23403G,G28881A,G28882A,G28883C EPI_ISL_6951092|2021-12-02|2020-02-10|B.1.1|China|Guangxi|Nanning|8|C241T,C3037T,C14408T,T20394C,A23403G,G28881A,G28882A,G28883C EPI_ISL_2737704|2021-06-30|2020-02-15|B.1.1|United Kingdom|England||8|C241T,C3037T,C14408T,A23403G,A27938G,G28881A,G28882A,G28883C EPI_ISL_2737705|2021-06-30|2020-02-16|B.1.1|United Kingdom|England||8|C241T,C3037T,C14408T,A23403G,A27938G,G28881A,G28882A,G28883C EPI_ISL_466615|2020-06-11|2020-02-16|B.1.1|United Kingdom|England||10|C241T,C3037T,C14408T,A23223G,A23403G,C25587T,T26483C,G28881A,G28882A,G28883C EPI_ISL_5248212|2021-10-17|2020-02-20|B.1.1|Ireland|Kerry||9|C241T,C313T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C,C29614T EPI_ISL_644823|2020-11-19|2020-02-22|B.1.1|USA|Washington||7|C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C EPI_ISL_423662|2020-04-12|2020-02-23|B.1.1|United Kingdom|England||7|C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C EPI_ISL_2737710|2021-06-30|2020-02-24|B.1.1|United Kingdom|England||8|C241T,C3037T,G11842A,C14408T,A23403G,G28881A,G28882A,G28883C EPI_ISL_437932|2020-05-12|2020-02-24|B.1.1|Austria|Ischgl||7|C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C EPI_ISL_460081|2020-06-05|2020-02-24|B.1.1|Italy|Lombardy||7|C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C EPI_ISL_412912|2020-02-28|2020-02-25|B.1.1|Germany|Baden-Wurttemberg||8|C241T,C3037T,G10265A,C14408T,A23403G,G28881A,G28882A,G28883C EPI_ISL_464184|2020-06-11|2020-02-25|B.1.1|United Kingdom|England||8|C241T,C3037T,C14408T,A23403G,A27938G,G28881A,G28882A,G28883C EPI_ISL_542235|2020-09-23|2020-02-25|B.1.1|Italy|Lombardy||7|C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C EPI_ISL_542237|2020-09-23|2020-02-25|B.1.1|Italy|Lombardy||7|C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C EPI_ISL_542238|2020-09-23|2020-02-25|B.1.1|Italy|Lombardy||7|C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C EPI_ISL_542410|2020-09-23|2020-02-25|B.1.1|Italy|Lombardy||7|C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C EPI_ISL_2731458|2021-06-30|2020-02-26|B.1.1|Brazil|Parana|Curitiba|7|C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C EPI_ISL_2737721|2021-06-30|2020-02-26|B.1.1|United Kingdom|England||7|C241T,C3037T,C14408T,A23403G,G28881A,G28882A,G28883C